Making eDNA data more visible, reusable, and impactful
Environmental DNA (eDNA) offers a powerful, non-invasive way to monitor biodiversity at both local and global scales. In December 2025 we used a bespoke open-source tool - the eDNA Bridge - to share over 750,000 eDNA observations from New Zealand with the global community.
New Zealand Ministry for the Environment
Data that is all around us
Every organism leaves traces of DNA in its environment—whether through skin cells, hair, pollen, or even slime. Environmental DNA (eDNA) sampling captures these traces, offering a powerful way to profile biodiversity across ecosystems. To support the reuse and sharing of New Zealand eDNA data, the Ministry for the Environment commissioned the development of an open-source software package, developed with the widely used programming language R.
This project would not have been possible without our collaborators from Wilderlab, Hawke's Bay Regional Council, the Department of Conservation and Earth Sciences New Zealand.

Maximising the impact of data
Publishing biodiversity records to centralized repositories like GBIF ensures data are FAIR (Findable, Accessible, Interoperable, and Reusable), and opportunities for conservation impact are maximised. The eDNA Bridge tool improves the accessibility and interoperability of New Zealand’s eDNA data and supports national, regional and international collaboration.
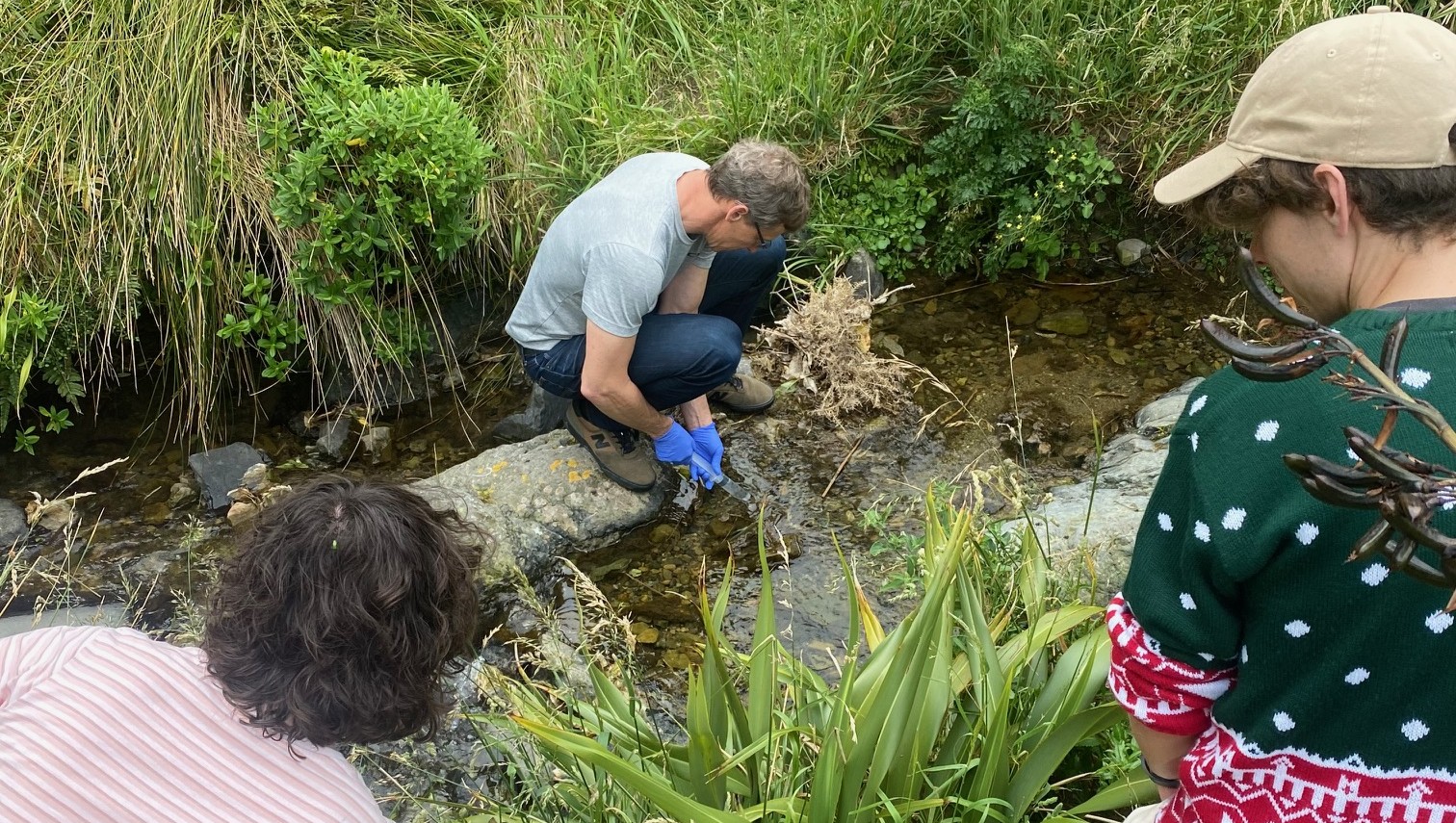

"Epi have been excellent to work with – both technically and socially. Sharing data requires trust, and I’ve really appreciated their ongoing and open communication to the data holders that volunteered to upload data."
“They worked with us (MfE), data owners, the primary commercial eDNA lab, GBIF NZ and GBIF International to understand the wider context and challenges in data sharing. I’m so excited to see their code enable automated upload of eDNA direct from the lab, enabling us to get the full value from New Zealand’s investment in eDNA data.”
Fiona Hodge
Senior Advisor (Science), Ministry for the Environment
Removing technical bottlenecks
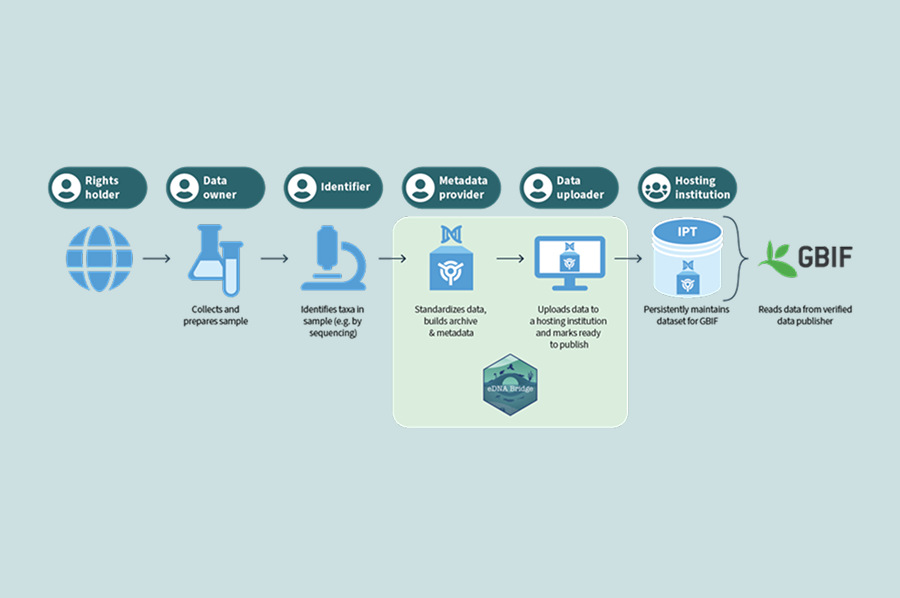
Several operational barriers limit data sharing, preventing the widespread dissemination and use of eDNA data. The tool addresses the most labour-intensive technical part of the data sharing process: the preparation, validation, and bulk submission of data to Global Biodiversity Information Facility (GBIF). The eDNA Bridge tool enables bulk upload of eDNA data to the global repository, where data can be accessed by anyone.
The package has already been adapted by Wilderlab, New Zealand’s pioneering eDNA laboratory for routine submission of results to GBIF, for consenting data owners.

Executed to support wide adoption
The eDNA Bridge is an open-source, extendable, R package that provides an end-to-end, scalable workflow from sequencing laboratory outputs through to publication on GBIF. The package is designed for flexible use, supporting both interactive, guided workflows and fully automated, scripted integration within laboratory or organisational pipelines. Improving data quality, consistency, and interoperability by producing GBIF-compliant Darwin Core Archives and supporting programmatic upload to Integrated Publishing Toolkit (IPT) servers. For ease of use the package includes detailed user guidance in the command line interface, high level functions to automate uploads with sensible default values and low-level functions that can be incorporated into custom R scripting workflows.
The eDNA Bridge can be downloaded for free from the GBIF GitHub page here.